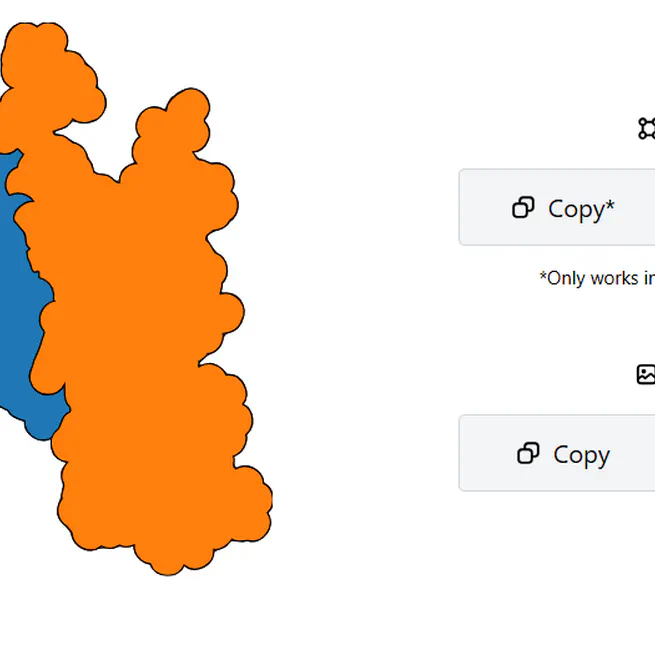
Takes as input a protein structure and vectorizes it in 3 styles: Flat, 3D or Goodsell.
Mar 25, 2025
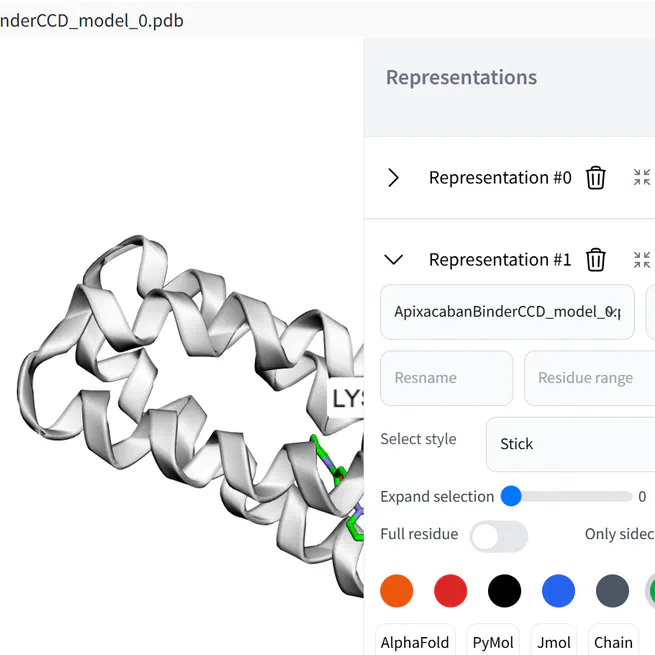
Gradio component to visualize 3D molecules using 3dmol.js. Allows to change representation on the fly.
Mar 25, 2025
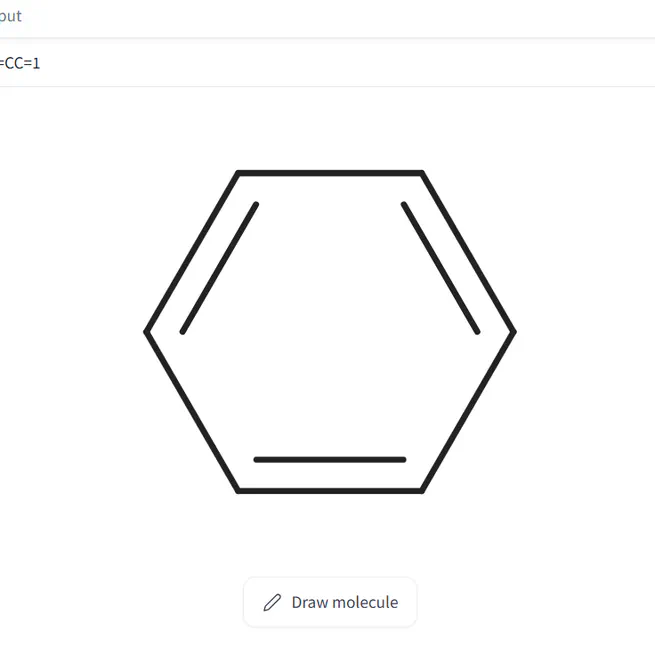
Gradio Molecule2D allows to visualize and draw molecules using the ketcher library.
Mar 25, 2025

Gradio component to set up cofolding models like AlphaFold3, RoseTTAfold-AllAtom, Chai1…
Mar 25, 2025
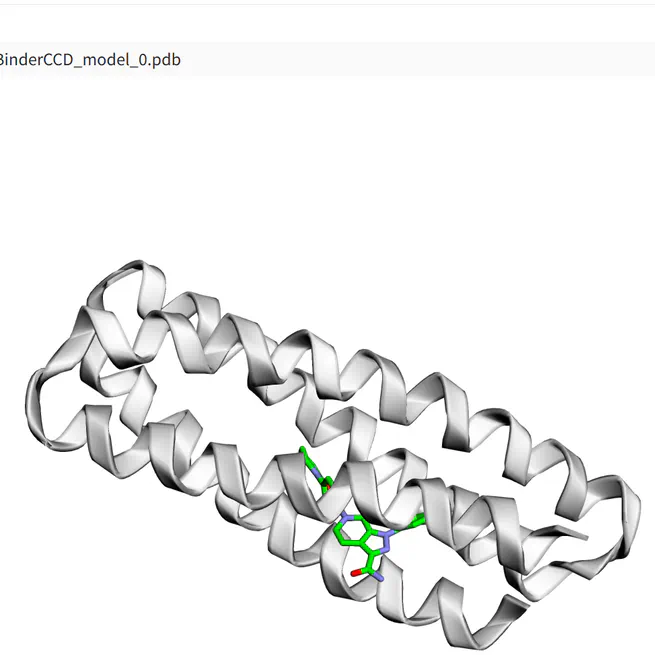
Interface for Boltz-1 model by Wohlwend et al built using gradio_cofoldinginput
Mar 25, 2025
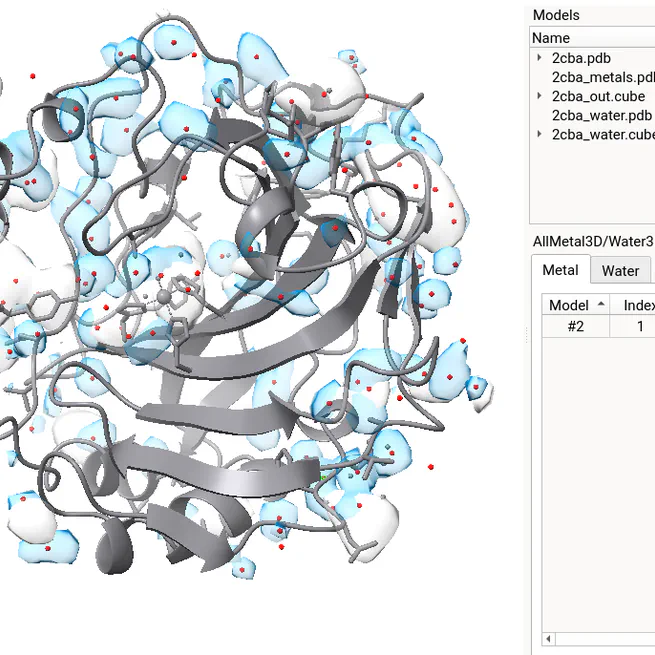
AllMetal3D allows to localize and classify metal-binding sites in proteins. Available as webapp and ChimeraX extension.
Mar 25, 2025
Selectively remove solutions from a private teacher repository using GitHub actions. Also allows removing cells tagged solution in a jupyter notebook.
Apr 1, 2021
Webapp for ProteinMPNN with advanced selection capabilities and refolding of designed sequences using AF2 in single sequence mode
Apr 1, 2021
Database of permissively licensed vector illustrations
Apr 1, 2021
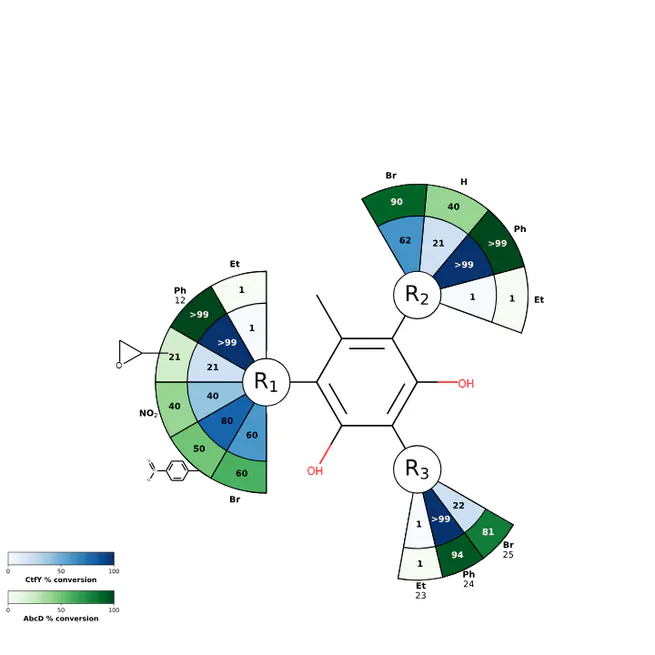
Visualize substrate scope on a molecular structure.
Jan 1, 2019